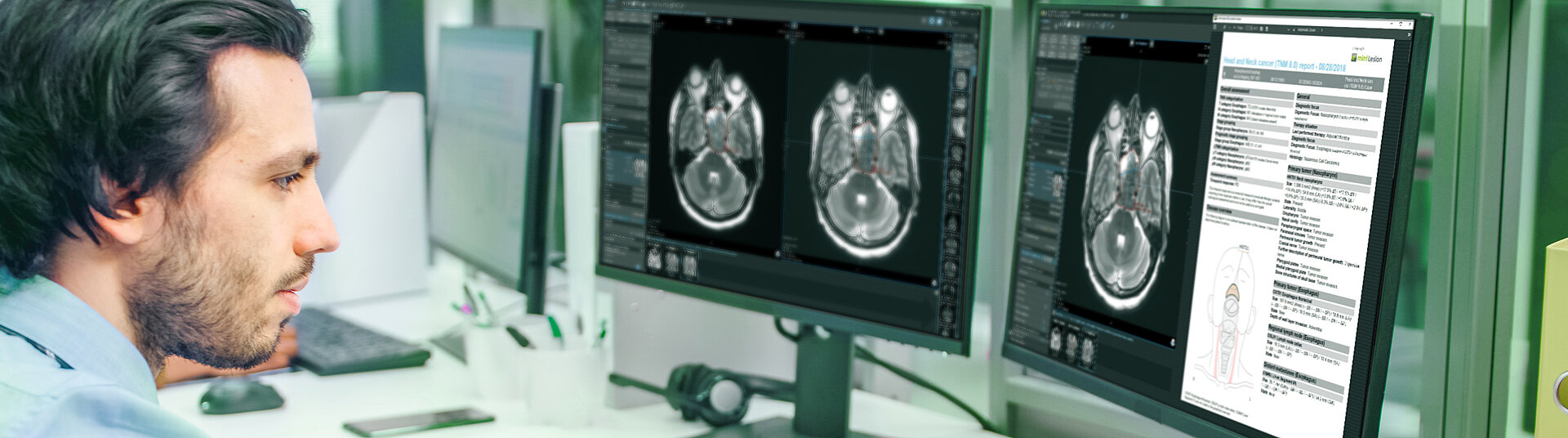
Transform Diagnostic Radiology with Structured Reporting and Data-Driven Intelligence
mint Lesion is a structured reporting software for oncological imaging with comprehensive reading templates, standardized data integration and analytics capabilities
Connect with us to discover how AI-powered, structured, data-driven reporting can function within your radiologic operations while positioning you for future innovation.
Guideline-Based Structured Assessment for Multi-Entity Diagnostics
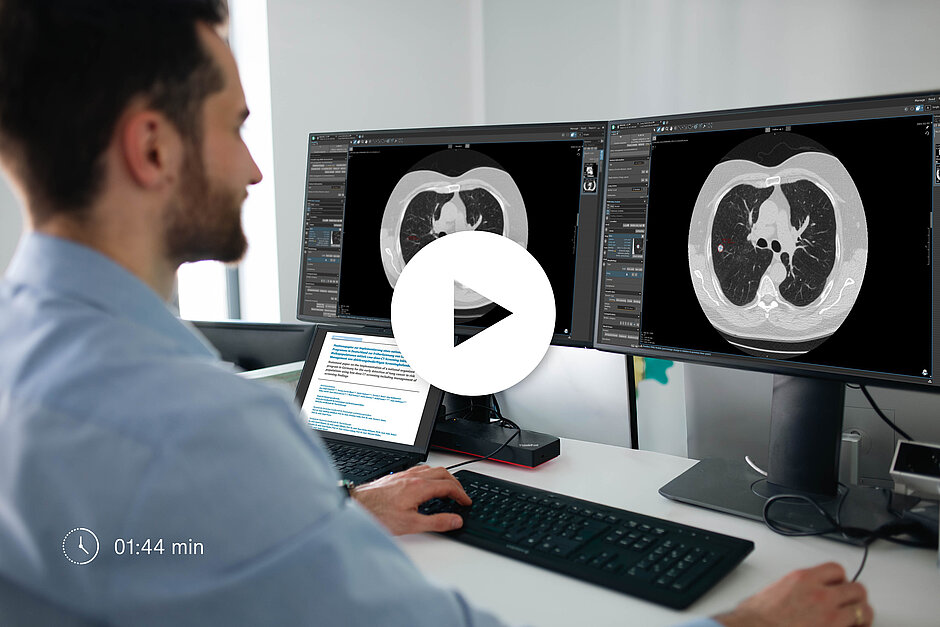
Comprehensive Support for Lung Cancer Screening and Nodule Management
- Standardized Scoring Systems: mint Lesion provides a guided workflow for the categorization of lung nodules according to the Lung-RADS 2022 framework, Brock model malignancy risk assessments, and ACR management recommendations.
- Configurable Reporting Templates: mint Lesion templates can be tailored to local or institutional needs – for example, adapting to Incidental Pulmonary Nodule (IPN) management protocols – while ensuring alignment with international guidelines.
- Automated Nodule Volumetry: mint Lesion integrates Third-party AI tools that support automated detection and characterization of nodules.
- Nodule Growth Tracking: mint Lesion automatically determines volume doubling time (VDT) based on the patient’s longitudinal data.
- Open 360° Ecosystem: mint Lesion offers a comprehensive end-to-end solution, streamlining the entire clinical pathway from initial participant enrollment to registry integration and supporting advanced clinical research initiatives.
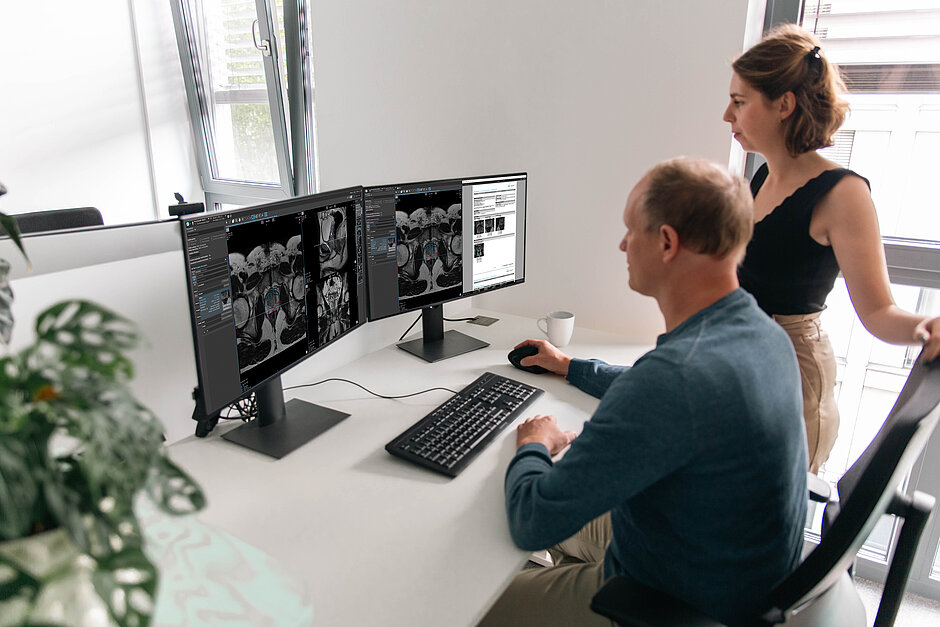

Structured Prostate MRI Evaluation using PI‑RADS v2.1
- PI-RADS v2.1 Compliance: mint Lesion provides a guided workflow based on the current PI-RADS classification, offering a structured interface for the targeted assessment of the peripheral and transition zones.
- Automated Segmentation: mint Lesion incorporates third-party AI software for automated prostate segmentation, volume calculation and lesion detection within T2w, DWI/ADC, and DCE sequences.
- PSA Density: By combining the calculated volume with the PSA value, mint Lesion automatically determines the PSA density.
- Targeted Biopsy Interoperability: To support guided biopsy procedures, mint Lesion exports findings as DICOM Segmentation Objects for direct integration into urological biopsy devices.
- PI-QUAL v2 Assessment: mint Lesion integrates the PI-QUAL v2 score into the prostate workflow, allowing radiologists to objectively evaluate and document the diagnostic quality of mpMRI scans within the structured report.
Multi-Entity Tumor Staging using TNM Classifications
Across various oncological entities, mint Lesion implements the TNM staging criteria. The software maps measured diameters and invasion depth directly to T, N, and M categories, ensuring that the structured report reflects the current international staging standards.
Customizable Reading Templates for Therapy Response Assessment
The structured reporting framework in mint Lesion allows for the configuration of reading templates ↗ that adapt to specific clinical requirements.
- Quantitative Measurement Tracking: mint Lesion records changes in findings (diameter/volume) and functional parameters (SUV/ADC) relative to baseline within a structured report.
- Minimum Size Validation: Templates enforce compliance with required minimum lesion sizes to ensure data consistency.
- Clinical Biomarker Integration: The platform allows for the side-by-side visualization of structured reporting metrics with clinical values such as PSA or ctDNA.
- Interactive Longitudinal Charts: mint Lesion generates dynamic charts based on structured reporting data to display correlations between tumor findings and clinical values.
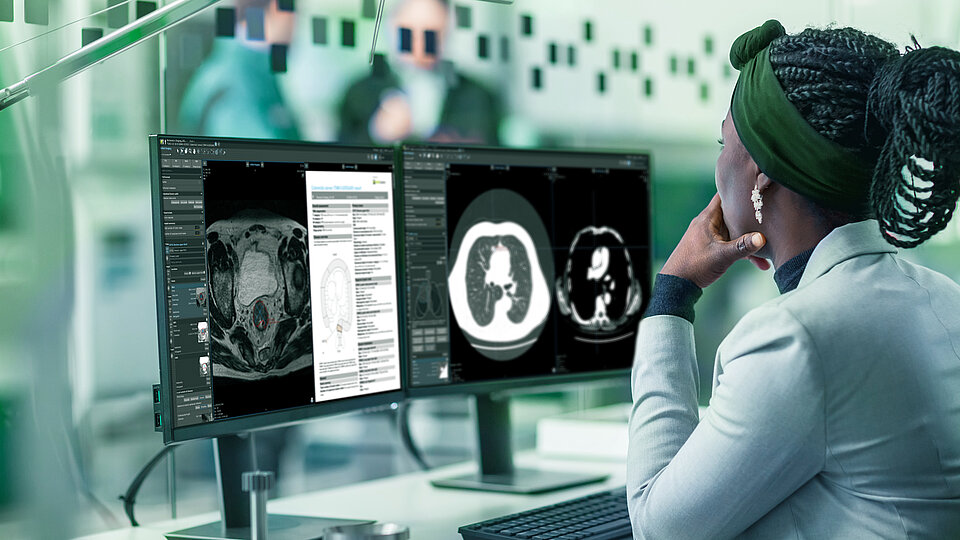
Rely on your cognitive assistant at eye level
Experience guided reporting
To ensure a complete and comprehensive report containing all information necessary for holistic therapy management...
Know the reason why
The uniform, intuitive user interface of mint Lesion is optimized for each clinical application scenario...
Speed up your work
mint Lesion automates time-consuming routine procedures, allowing you to focus on the images...
Experience guided reporting
To ensure a complete and comprehensive report containing all information necessary for holistic therapy management, mint Lesion™ walks you through the reporting process in an interactive, dynamic dialogue. The software knows each patient’s case and history as well as the appropriate diagnostic criteria and guidelines. It asks you the right questions at the right time, and reliably prepares the results for you.
Know the reason why
The uniform, intuitive user interface of mint Lesion™ is optimized for each clinical application scenario. The software knows all the intricacies of integrated guidelines, performs conformity checks throughout your reading process, and notifies you of any discrepancies. For each automatically generated evaluation result, such as the T-stage in context of a TNM cancer classification, the software provides a comprehensive explanation.
Speed up your work
mint Lesion™ automates time-consuming work steps of radiological reading, allowing you to focus on the images. With advanced tools for the different modalities and streamlined workflows for screening, staging, and tumor response evaluation, mint Lesion™ saves you time and effort. Its AI-driven workflow assistance, as well as automated image correlation, lesion matching and personalized work lists accelerate your reading process substantially.
You can start mint Lesion™ directly from your RIS / PACS while preserving the working context. Relevant image data will be automatically synchronized with your PACS, and results can be integrated into your RIS report or EMR/HIS system.
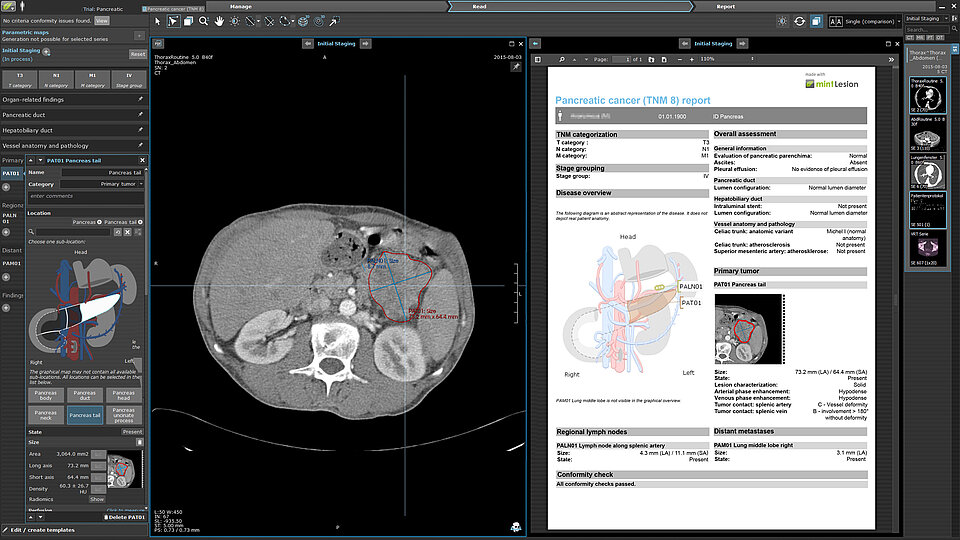
Be always at the forefront
Promote data-driven radiology and integrated diagnostics
With mint Lesion, you transform radiological images into a digital stream of quantitative diagnostic data...
Gain new knowledge from images
mint Lesion provides you with longitudinally connected data beyond size and location of a tumor...
Apply up-to-date guidelines
Integrated guidelines are updated regularly to keep you abreast of the latest developments...
Promote data-driven radiology and integrated diagnostics
With mint Lesion™, you transform radiological images into a digital stream of highly structured and quantitative diagnostic data. Along with gaining a wealth of information from images, radiologists strengthen their position as the knowledge holders of patient- and disease-related specifics and drive information exchange between medical disciplines. mint Lesion™ facilitates your systematic data integration with other sources of diagnostic data – e. g. from pathology and clinical laboratory that serve as the basis for clinical decisions.
Gain new knowledge from images
mint Lesion™ provides you with longitudinally connected data beyond size and location of a tumor. In the background of your daily reads, you can extract and export 1st and 2nd order radiomic features as well as tumor growth rates (g-values) on a single patient- or patient cohort level. mint Lesion™ converts images into mineable data. Subsequent analysis of these data can improve diagnostic accuracy and prognostic assessment, and increase predictive power for decision support tools.
Apply up-to-date guidelines
Integrated guidelines are updated regularly to keep you abreast of the latest developments in medicine. You can read up on new reading templates directly in mint Lesion™ and learn how to apply them in a complete and formally correct way during the guided reading process.

Enhance your report quality and communication
Generate structured, high-quality reports
The collected data with their clinical context are clearly structured and dynamically integrated into...
Improve your interdisciplinary communication
Reports generated with mint Lesion are clearly and consistently structured to make complex information easily understandable...
Help your patients to understand
Next to containing distinct information, mint Lesion reports include snapshots of findings and graphs indicating...
Generate structured, high-quality reports
The collected data with their clinical context are clearly structured and dynamically integrated into the complete and easy-to-understand reports. With a standardized level of detail, structure and terminology, the generated reports provide a solid basis for treatment decisions, e.g. at tumor board meetings.
Snapshots of findings are integrated directly into the report, and their locations are automatically indicated in the accompanying anatomical diagrams.
Reports are available in DICOM, HL7, PDF, XML and Office formats that can be easily transferred into your RIS, PACS and HIS.
Improve your interdisciplinary communication
Reports generated with mint Lesion™ are clearly and consistently structured to make complex information easily understandable. Your colleagues from other disciplines will easily find the information that is relevant to them, which leads to fewer queries and enables a cooperative approach.
Help your patients to understand
Next to containing distinct information, mint Lesion™ reports include snapshots of findings and graphs indicating various parameters such as tumor volume or growth over time. These visual depictions can help patients better understand the course of their disease and facilitate the way you and your colleagues communicate with your patients.
What our users say about mint Lesion
Since 2013, mint Lesion has provided dedicated reporting profiles for oncological radiological diagnostics. The software tracks measurements with their semantic context, enabling a reading process that directly links findings to imaging data.
The mint approach to structured reporting combines imaging and clinical data to support completeness, reproducibility, and clear interdisciplinary communication.
Standardized data can be used across medical departments and compared within patient groups or across populations. This supports treatment recommendations, quality assurance processes and clinical decision-making.
The structured data format provides a foundation for big data analytics and AI applications as well as improved or new guidelines, helping users adapt to evolving healthcare technology while maintaining consistent reporting standards.



