A recent study [1] conducted by researchers at University Hospital Ulm explored the role of sarcopenia in patients with esophageal or gastric cancer. The main objective of this study was to establish a framework for categorizing the impact of sarcopenia on the prognosis of individuals diagnosed with these specific cancer types. In addition, the researchers investigated the relevance of CT radiomics for both diagnosis of sarcopenia and prognosis of tumor progression.
Data from 83 patients who underwent contrast-enhanced CT scans between 2015 and 2019 were analyzed. The prevalence of sarcopenia, defined by the psoas muscle index (PMI), was examined at three time points: initial diagnosis (t1), after neoadjuvant chemotherapy (t2), and one year post-surgery or post-chemotherapy (t3). The PMI values demonstrated a significant decrease from t1 to t2 and from t2 to t3. Disease progression was monitored according to the response evaluation criteria in solid tumors (RECIST 1.1). On average, patients with sarcopenia did not experience progressive disease (PD) more frequently than non-sarcopenic patients at any time point.
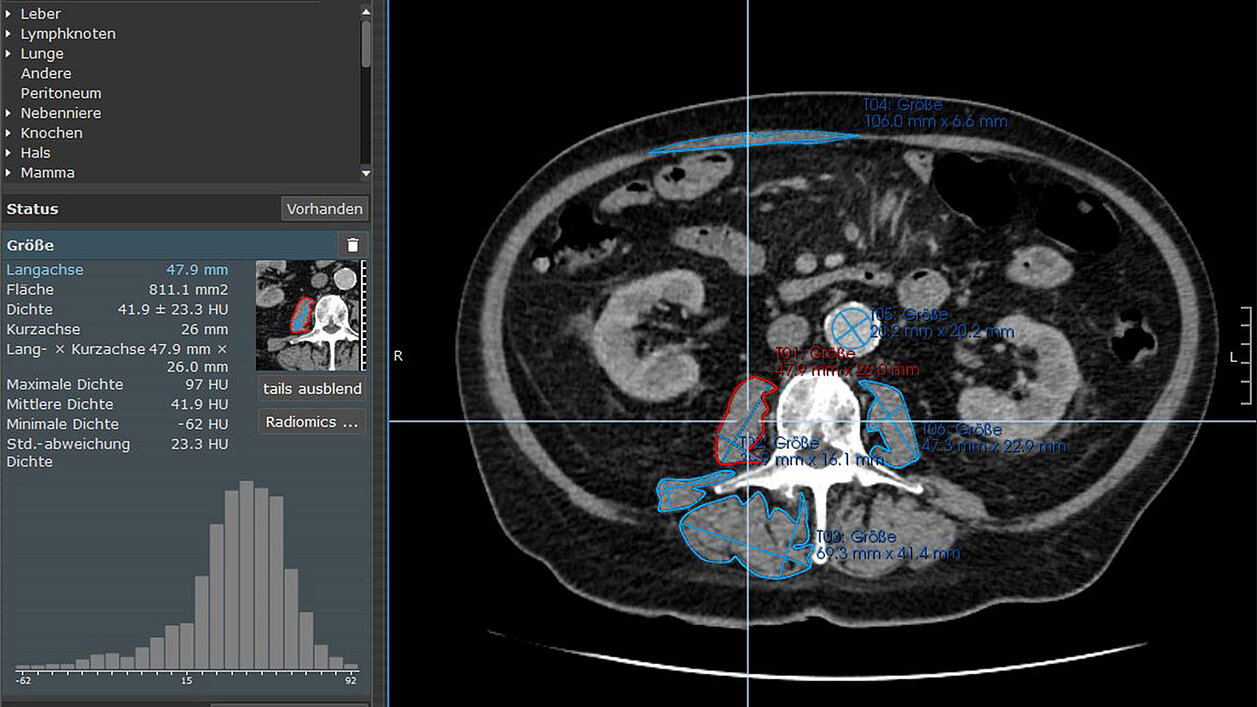
mint Lesion™ was used for skeletal muscle segmentation and radiomic feature extraction. The researchers segmented the psoas major muscle, erector spinae muscle, and quadratus lumborum muscle and extracted a comprehensive set of 85 radiomic features from each analyzed skeletal muscle.
The study utilized these radiomic features to provide a more in-depth assessment of skeletal muscles. The features were categorized into different types:
- Shape features, which describe the geometry and morphology of the segmented skeletal muscle - e.g., volume, sphericity, and other shape-related parameters;
- First-order features, which depict the distribution of voxel intensities within the defined region of interest (ROI) - e.g., mean intensity, median intensity, and standard deviation;
- Higher-order features, which go beyond simple statistics and involve more complex analyses of voxel values by reflecting the arrangement and relationships between voxel intensities within the ROI – e.g., cluster tendency, difference entropy, and sum of entropy.
“In comparison to classical muscle segmentation, radiomic features offer more information for qualitative evaluation, providing more potential for evaluating qualitative patterns,” stated the researchers. “Given the quantitative and additionally qualitative differences in skeletal muscles of patients with tumor diagnosis and muscle loss, radiomics might be helpful to detect sarcopenia and predict tumor progression.”
The study aimed to assess the diagnostic and prognostic value of extracted radiomic features in combination with machine learning models, specifically Decision Tree (DT), K-nearest neighbors (KNN), and Random Forest (RF). For sarcopenia prediction, the RF model demonstrated high accuracy (0.93) and area under the curve (AUC) values (0.97) for the psoas major muscle at the initial diagnosis (t1). This suggests a strong potential for CT radiomics and machine learning to assist in the early detection of sarcopenia and prediction of tumor progression in oncological patients.
While the association between PMI-diagnosed sarcopenia and disease progression did not reach statistical significance, the study underscores the potential of CT radiomics and machine learning in clinical practice. It emphasizes the need for standardized approaches and highlights the role of innovative methods in enhancing diagnostic and prognostic processes in oncological imaging.
Read the original publication here.
[1] Vogele D, Mueller T, Wolf D, Otto S, Manoj S, Goetz M, Ettrich TJ, Beer M. Applicability of the CT Radiomics of Skeletal Muscle and Machine Learning for the Detection of Sarcopenia and Prognostic Assessment of Disease Progression in Patients with Gastric and Esophageal Tumors. Diagnostics 2024.